Configuration Networks
Configuration Networks, Protein Folding Landscapes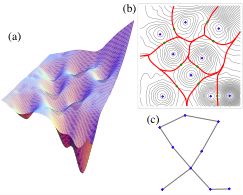
- Focus:
- Robot arm model: build constraied version, characterise constrains in hyper-lattice space, include forces andcharacterise gradient networks and landscapes
- Model configuration spaces: study the effect of topology, constrain statistics and energy-topology interplay
- Random geometric lattice: check landscape properties, prove that minimum bias is enough, solve analythically
- Real systems: measure relevant properties of many real configuration spaces (Rao, Doye, Scala, ect.)
- Summary: Protein Folding Networks and Levinthal’s Paradox (LA-UR-05-3525)
- Bibliography:
- C. Levinthal, How to Fold Graciously, Mossbauer Spectroscopy in Biological Systems (proceedin
gs), Allerton House, Monticello, Illi
nois, Eds: J. T. P. DeBrunner and E. Munck, University of Illinois Press, 22 (1969);
- D.B. Wetlaufer, Nucleation, Rapid Folding, and Globular Intrachain Regions in Proteins, Proc. Natl. Acad. Sci. USA 70, 691 (1973);
- D. Wales, M.A. Miller, T.R. Walsh, Archetipal Energy Landscapes, Nature 394, 758 (1998);
- A. Scala, L. A. Nunes Amaral and M. Barthélémy, Small-world networks and the conformation space of a short lattice polymer chain, Europhys. Lett. 55, 594 (2001);
- F. Rao and A. Caflisch, The protein folding network, J. Molec. Biol. 342, 299 (2002);
- J.P.K. Doye and C.P. Massen, Evolution of the potential energy surface with size for Lennard-Jones clusters, J. Chem. Phys. 111, 8417 (1999);
- J.P.K. Doye and C.P. Massen, Characterizing the network topology of the energy landscapes of atomic clusters, cond-mat/0411144, (2004);
- Z. Toroczkai, K.E. Bassler, Jamming is limited in scale-free systems, Nature 428, 716 (2004);
- Z. Toroczkai, B. Kozma, K.E. Bassler, N.W. Hengartner and G. Korniss, Gradient Networks,
- J. Dall and M. Christensen, Random geometric graphs, Phys. Rev. E, 66, 016121 (2002);

